|
Applications of MR Image Segmentation
Tomi Heinonena, Prasun Dastidarb, Harry Freyc, Hannu Eskolaa
a Ragnar Granit Institute, Tampere University of Technology, Tampere, Finland
b Department of Radiology, Tampere University Hospital, Tampere, Finland
c Department of Neurology, Tampere University Hospital, Tampere, Finland
Abstract.After the introduction of digital imaging devices in medicine computerized tissue recognition
and classification (i.e., segmentation) have become important in research and clinical applications. Segmented
data can be applied among numerous research fields including volumetric analysis of particular tissues and structures,
construction of anatomical models, three-dimensional (3D) visualization, and multimodal visualization, hence making
segmentation essential in modern image analysis. In this research project several PC based software were developed
in order to segment medical images, to visualize raw and segmented images in 3D, and to produce EEG brain maps
in which MR images and EEG signals were integrated. The software package was tested and validated in numerous clinical
research projects in hospital environment.
1. Introduction
During the last three decades computerized segmentation techniques have been studied among numerous scientific
fields including, e.g., pattern recognition, computer vision systems, and medical imaging. In general, most of
the basic methods can be utilized among all segmentation applications. Various segmentation techniques can be classified
using two criteria: operation principle or the level of interactivity. The operation principle can be classified,
for example, as boundary based technique, region based technique, or statistical technique. The level of interactivity
in turn can be classified as manual, semiautomatic, or automatic. Different techniques can be integrated hence
making the accurate classification difficult. Both manual and automatic techniques have their advantages and drawbacks;
Automatic methods are sensitive to noise and unexpected situations leading to errors which will not be corrected,
but the results are highly reproducible. Manual and semi-automatic methods are not as sensitive to noise, but reproducibility
is usually poor because the operator can affect to the accuracy of results considerably. The quantitative comparison
of different segmentation algorithms is very difficult because numerous issues must be taken into account. Still
several statistical approaches have been suggested to evaluate the efficiency, advantages, and limitations of different
algorithms [1]. Segmentation appears to be a key issue in modern medical image analysis enabling numerous clinical
applications, such as three-dimensional (3D) visualization of separate neural structures in magnetic resonance
(MR) images, reconstruction of anatomical models, and volumetric analysis of MR images (Fig. 1).
Segmented neural structures can be applied in the reconstruction of piecewise homogenous digital head models.
Segmented MR images can be applied directly in forming conductivity models consisting of volume elements, such
as finite difference element models (FDM) [2]. Because bone tissue has a relatively low conductivity, it affects
strongly the EEG amplitude measured from the scalp. Also the variation of scalp thickness can cause 20-40% variation
in EEG amplitude [3,4]. A reasonable head model requires at least the segmentation of scalp, skull, CSF, gray matter,
and white matter of the brain. The models can be utilized, e.g., in simulations enabling electrical modeling of
the brain activity [4,5]. Head modeling enables numerous applications, such as the detection of epileptic foci
and other sources [6,7,8]. Head models can be applied also in other simulations, such as computing the absorption
of electromagnetic radiation [9].
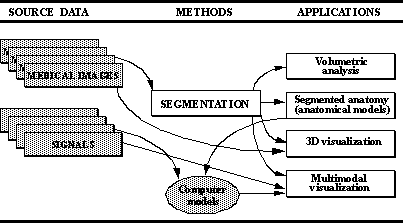
Figure 1. Segmentation and it's applications. Segmented anatomical structures can be applied
in volumetric analysis and reconstruction of head models. Further, additional information, such as measured or
simulated signals can be integrated with 3D presentation in order to form multimodal representations.
Due to increasing accuracy of MR images and segmentation it is possible to define non-invasively the volumes
of different organs, neural structures, and pathological lesions. In addition, various neurological conditions,
such as brain atrophy, edema associated with various disorders, and brain hematomas can be estimated based on volumetric
analysis. Multiple Sclerosis (MS) has been widely studied utilizing volumetry [10-17]. The volume of MS plaques
and increasing cerebral atrophy have been measured and correlated to clinically measured signs of the disease.
Other common volumetric applications are the analysis of different tumors [18,19], brain infarcts [20,21], and
neurodegenerative diseases [22,23]. The measured volumes of particular neural structures and pathologies can be
utilized in determining appropriate treatment. Further, a set of consecutive MR studies together with volumetry
enable an efficient follow-up of disease progression. Such technique can be applied e.g., in the treatment of MS
and the radiation therapy of tumors.
MR images can be displayed three-dimensionally enabling efficient analysis of MRI data; anatomical
structures are easier to render from 3D images than from conventional 2D slices. 3D graphics have been applied
among clinical medicine in order to plan operations [24] and visualizing tumors and other anomalies [25]. Three-dimensional
visualization principles of MR images can be divided into surface rendering or volume rendering. Although volume
rendering is capable of producing 3D images directly, it is often utilized in order to produce surfaces, hence
the visualization can be understood as a two step procedure consisting of surface reconstruction and rendering.
Surface reconstruction can be classified into several different techniques, based on polygon surfaces, parametric
surfaces, and volume rendering [26]. The selection of appropriate reconstruction technique depends on the type
of source data; raw MR images can be displayed utilizing thresholding and volume rendering techniques, boundary
segmented structures can be displayed using polygon or parametric surfaces, etc. Rendering techniques involve different
shading and displaying [26-29] methods.
One interesting application of 3D visualization is the multimodal imaging, in which different
types of information are integrated. Conventionally, multimodal imaging is associated with image fusion (e.g.,
combination of MRI and CT, SPECT, etc [30]. Nevertheless, completely different information modalities can be combined,
such as digital EEG and MRI data improving spatial resolution of EEG, and enabling comparison of EEG and MRI findings
[31-33]. The integration of EEG and MRI consists of several stages, including image segmentation, definition of
electrode locations, and the reconstruction of 3D surface. Thresholding is usually applied to segment the scalp.
Nevertheless, other segmentation techniques can also be utilized especially when other tissues than scalp are required.
The definition of electrode locations can be solved by two different techniques: Electrode locations can be marked
on MR images by using e.g., T1 MRI and vitamin E capsules [4] which appear excellently in MR images. However, when
the number of electrodes is high (e.g., 64 or 128), the marking becomes complex. Another approach is to mark or
detect only several landmarks, such as inion, nasion, and pre-auricular points, enabling computerized calculation
of electrode locations [34,4].
The 3D surface is usually reconstructed using triangulation [4], parametric surfaces [35-37], or volume rendering
[38]. When triangles or parametric surfaces are applied, various techniques can be applied to code and display
colors representing measured EEG data on surface elements, and in addition, different interpolation techniques
have been studied involving nearest electrodes [35,39]. If direct volume rendering is utilized instead of surface
rendering, the EEG data can be coded directly to the surface voxels as colors. One approach is to code EEG data
to another 3D surface consisting of triangles or parametric surfaces, and match it with the volume rendered surface
[38].
2. Material and Methods
During this study, several segmentation and visualization algorithms were developed and integrated into three
separate software (Fig. 2): The IARD [40] segmentation algorithm was developed to produce head models from MR images,
consisting of the scalp, skull, CSF, gray matter, and white matter. An interactive user interface together with
additional segmentation techniques (Anatomatic) was developed to enable interaction and volumetric analysis [41].
The volumetric accuracy of the developed algorithms and software was tested in the analysis of MS and brain infarct
[42]. A 3D visualization algorithms and software was developed (Medimag) to represent segmented structures and
multimodal images in 3D [43]. Also a technique able to represent EEG data and patient MR images (3DEEG) was developed
[38]. The developed segmentation and visualization software were applied in a clinical study on multiple sclerosis
[44].
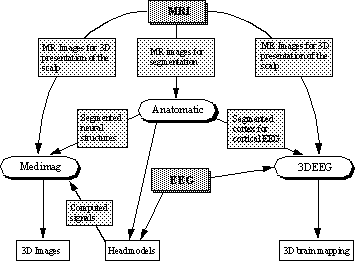
Figure 2. The developed software and data flow. Anatomatic software together with IARD segmentation
algorithm form an efficient segmentation tool. Medimag software is capable of displaying raw and segmented MR images
in 3D. 3DEEG software is able to visualize EEG and MRI data.
2.1 Segmentation Algorithm
Conventional image processing techniques were applied in the segmentation process. The basic tool in all operation
is the IARD (Image enhancement, Amplitude segmentation, Region growing, Decision tree) segmentation procedure [40].
Even if the IARD method includes the use of several existing image processing techniques, it is itself a new technique
which can be applied excellently in medical image segmentation. The first step in IARD technique consists of three
amplitude segmentation operations applying appropriate threshold coefficients. The coefficients can be obtained
directly from the image histogram of defined manually. The three amplitude segmented images are presented simultaneously
on the screen and can be modified using particular tools such as line or rectangle drawing and fill with connectivity
(2D Region growing). Various colors represent different tissues. When images appear appropriate they can be integrated
using several decision rules. Segmentation tools consist of image filtering, amplitude segmentation, region growing,
line and rectangle drawing, free hand drawing, and drawing with a small and large pen, etc.
Anatomatic Software
The segmentation software called Anatomatic was developed to 32bit Windows™ system (Win95™
and WinNT™) using ANSI C and C++ programming languages. It requires at least 16MB of RAM, minimum 16bit
colors and the minimum resolution of 1024X768 pixels. However, it is recommended to use 32MB RAM, 24bit colors,
and 1280X1024 resolution. Anatomatic can open, produce, save, and display medical images and segmented data. Further,
the data can be observed from different projection planes (sagittal, axial, and coronal) and volumetric analysis
are reported automatically.
The user interface of Anatomatic appears similar to a bitmap drawing program and is presented in Fig 3. First
the segmentation technique (IARD, manual, reloading, and lesion tracking) is selected. Then an image stack of MR
or CT images is opened and displayed on the screen. Current slice is defined and appropriate threshold coefficients
are selected. Thereafter a segmentation tool and a color representing a tissue are selected and the images are
modified. After the result appears realistic the user stores the segmented data and begins to process the next
image. The above mentioned steps are repeated as many times as there are images in the data set. However, one image
can be processed as many times as required, e.g., when several tissues are detected. If the appearance of images
is too small the window can be zoomed.
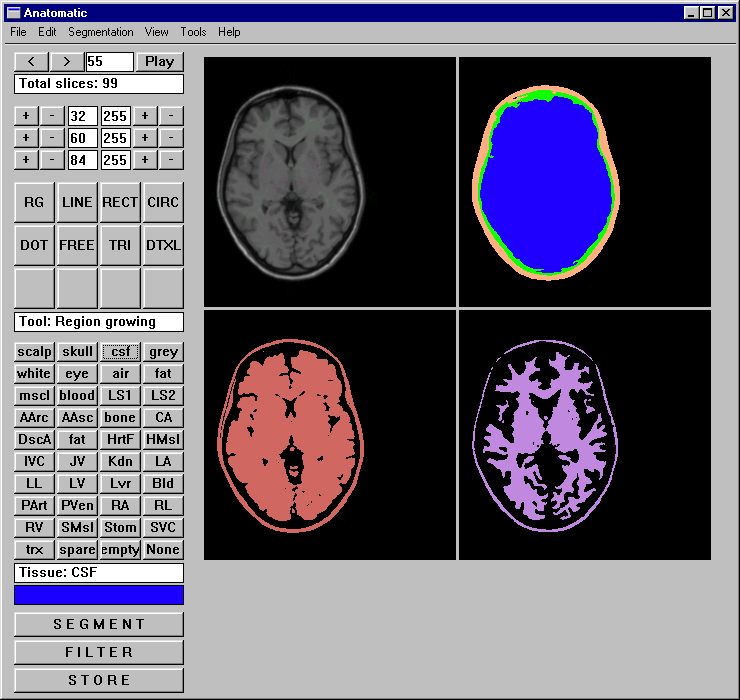 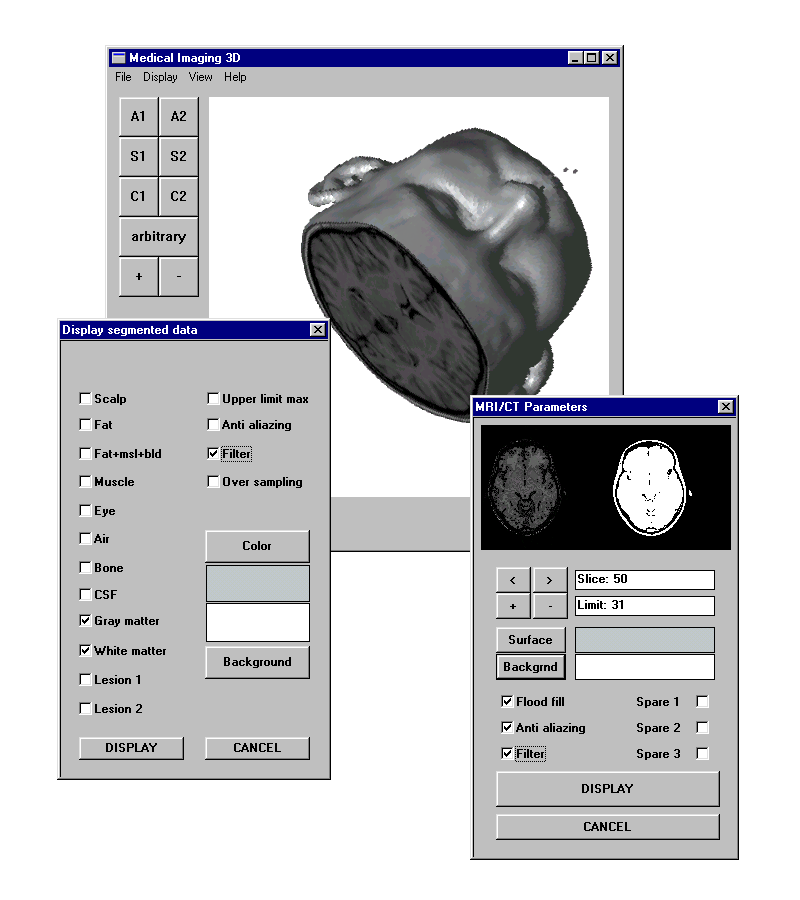
Figure 3. The user interfaces of Anatomatic and Medimag software. Anatomatic can be applied
in various segmentation procedures and applies IARD algorithm. Medimag is capable in visualizing raw medical images
(applying thresholding) and segmented data.
Volumetric Analysis
Because the physical dimensions of voxels are known it is possible to calculate the volumes of segmented structures.
The volumetric reports can be displayed any time during segmentation. The report include slice dependent information
about the number of voxels and volumes of different tissues together with the same information about the entire
image set. In addition, some parameters are computed automatically, such as brain volume (grey + white matter)
and relative atrophy (CSF / brain).
2.2 Visualization of MR Images
In order to present raw and preprocessed medical images in 3D a special computer software called Medimag was
developed. It was tested by using various test materials including several T1-weighted MR image sets of the head
(250 X 250 pixels, 100 slices, produced by a General Electric 0.5T MRI unit) and CT image sets of the skull (512
X 512 pixels, 100 slices, produced by a General Electric Pro Speed PLUS scanner). Also multimodal images were created
by combining simulated 3D electric fields and MR images. Simulations were computed using a previously reported
software developed at Ragnar Granit Institute for epileptic studies [2].
3D Visualization Algorithm
The developed visualization algorithm includes two separate implementations in order to detect a 3D surface.
The simpler one, called Simple Polygon Method (SPM) applies the basic Ray Casting theorem [27,28]: A 3D matrix
(e.g., MRI data) representing the modeled object is scanned from all six sides in order to find surface voxels.
The detected voxels are thereafter stored in a data structure for further processing. SPM operates perfectly when
the modeled object is a convex. However, cavities and complex structures cannot be detected accurately and therefore
another surface scanning technique called Flood Filling Method (FFM) was implemented. The basic conception of FFM
involves a non-recursive 3D flood filling [29] in order to detect and mark all the "empty" voxels outside
the object. FFM does not require additional memory for surface generation because it utilizes the same 3D matrix
where the voxel data is stored. The algorithm saves memory by using the voxel values under nine for its own computation
(limits the original MR/CT voxel values to the range of 10 to 255). Thereafter all the voxels adjacent to "empty"
voxels represent surface voxels. The criteria for determining voxels representing a particular object was based
on threshold coefficients.
The next step is to compute surface normal vectors for each surface voxel. That was implemented by applying
the "center of gravity" (CG) of a group of surface voxels; A voxel is surrounded by a small hollow sphere
(radius = 1 .. 5 voxels) and the CG is computed for this sphere. A vector drawn from CG to the voxel represented
the surface normal. The accuracy of the normal vector calculation depends on the size of the sphere and also on
some additional parameters. As a simple rule, large continuous surfaces can be processed using a large sphere and
small complex surfaces using a small sphere. However, high frequency noise on a continuous surface can be removed
using a large sphere.
Visualization of the surface voxels was implemented simply by applying Phong Shaded [28,29] polygons. Z-buffering
[29] was used in the process as a back-face removal algorithm. It also enabled the integration of two different
3D surfaces for presenting some segmented structures located partially within each other. The piecewise definition
of the surface color and transparency enables the visualization of arbitrary multimodal images.
Medimag Software
Medimag software was developed to utilize the developed 3D_object library. It involves the visualization of
original MR/CT images, produced 3D images, projection plane definitions, definitions of threshold coefficients,
etc. Medimag is capable of processing raw and RLE-coded segmented medical images. The appearance and operation
of the software are presented in Fig. 3. Medimag uses a simple data format to open MR/CT image sets. Any digital
image format can be easily converted to this format consisting of a header and data. The header contains image
stack size (x-, y-, and z-dimensions), physical dimensions, imaging techniques and projection planes. Image data
itself is stored as a 3D matrix using 8 bits per voxel.
Three-Dimensional EEG Brain Mapping
The process of displaying 3D brain maps consists of several stages. First electrode locations are approximated
or alternatively detected from MR images. E.g., in MR imaging such locations can be marked using particular oil
capsules. We have earlier developed a program to detect marked electrodes from MR image stack [2]. However, if
markings are not available there are various techniques to compute the locations [4]. First, some anatomical locations
such as nasion, inion, and pre-auricular nodes are defined. That enables the computation of the plane perpendicular
to the crown of the head. Thereafter the electrode locations can be calculated using analytical geometry. It can
be implemented e.g., by a set of lines following the surface of the head. Due to the interactive character of the
developed program, the user can select starting points directly from MR image stack using a mouse controlled cursor.
Thereafter the program defines automatically the electrode locations.
The next step is to generate triangles by using the electrodes as apices. That can be implemented, for example,
by applying heuristic triangulation methods [29]. However, when the number of electrodes is relatively small (e.g.,
10-20 system), pre-defined and optimized triangle mesh can be written into the program code. Virtual electrodes
are generated between the real electrodes thus increasing the number of triangles. That feature improves the shape
of the generated triangle mesh for modeling the head better. The potentials of the virtual electrodes are computed
using weighted averages of the surrounding electrodes.
For displaying EEG potentials we designed several color maps, such as frozen/heated object spectrum, RGB spectrum,
Blue-White-Red spectrum, etc. The frozen/heated object spectrum seems most natural; The spectrum changes from dark
blue(negative) to white(neutral) and then to dark red (positive). When an EEG signal sample is linked to the color
map it is possible to display the entire potential field using colors. For this purpose we applied Gouraud shading
[27] so that the colors corresponding to potentials in triangle apices were interpolated over the triangles. Such
method enables a smooth linear interpolation between electrodes and the result is an image representing electric
fields on the surface of the head.
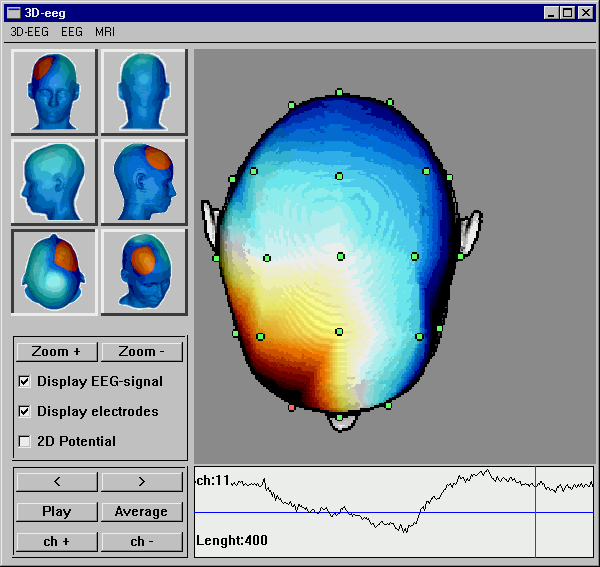 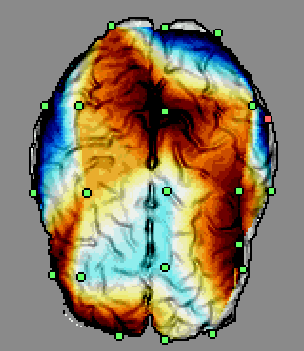
Figure 4. The user interface of 3DEEG software. Several projection planes can be selected.
The measured EEG signal can also be displayed as a smooth animation. The software is also capable in displaying
cortical EEG distributions.
In order to develop the software towards 3D effects, the methods of image fusion and ray casting
[27,28] were applied. Ray casting is a common method that can be applied in 3D visualization of any voxel based
data, especially in medical imaging. Our implementation of ray casting begins by computing the Z-buffer for a MRI
set. The Z-buffer is a 2D matrix representing the surface of a 3D object (e.g., the brain) located inside a 3D
matrix (MRI set). Because the Z-buffer is a 2D representation of 3D data, it can be obtained for one projection
plane at a time. The next stage is to smooth the Z-buffer using averaging, and then compute a surface gradient
which is a 2D matrix representing the shape of the 3D surface. When applied to MR image of the head the result
is a 2D matrix representing the surface gradient of the patient head. Several shading methods, such as Gouraud
[27] and Phong [27] can be utilized to display the matrix as a 3D bitmap. When this 3D bitmap and the 2D bitmap
representing electric fields are combined (image fusion) the result is a 3D image presenting electric fields on
the surface of the patient head. The image fusion can be implemented, e.g., by applying transparency.
The 3D-EEG program was developed for 32bit Windows operating systems (Win95, Win98 and WinNT) applying graphical
user interface enabling intuitive and efficient operation. The minimum requirements for the hardware are 16MB of
RAM and 16bit color display with the resolution of 800 X 600 pixels. An example of the program's appearance can
be seen in Fig 4.
3. Results
MR images of five epileptic patients were segmented using manual and semi-automatic IARD techniques to five
different tissues: scalp, skull, CSF, gray matter, and white matter. The results were applied in resistive head
models and simulations. It was realized that semiautomatic IARD operates efficiently only in the region between
the top of the head and eyes. Other regions can also be segmented but requires more interaction. The average processing
time depends on the quality of images and what tissues are segmented and varies from 10 to 120 seconds per slice.
Although T1 MR images were applied in segmentation (CSF and bone cannot be separated), the IARD algorithm enabled
approximation of the interior of the skull, based on prior knowledge of anatomical structures and decision trees.
It was realized that semiautomatic IARD is capable of segmenting the interior of the skull with 65% accuracy according
to the criteria that there should be 2-5 mm of CSF between the brain and skull. The accuracy can be increased by
using lowpass filtering and by increasing interaction. Due to interactive character of the IARD algorithm, the
person who controls the segmentation procedure should be trained and elaborate in order to produce accurate results.
Anatomatic software was tested with varying test material and segmentation techniques: Six epileptic patient
images (100 axial T1 MRI slices per patient covering the entire head) were segmented into scalp, skull, grey matter,
and white matter. The segmentation required 2-3 hours of work for each patient, and the results were anatomically
realistic, including the interior of the skull. A MRI set of human head (120 axial T1 slices) was segmented into
11 separate tissues including scalp, fat, muscle, bone, etc. Such segmentation procedure required 24 hours of processing.
Visible Human Man (VHM) images (reduced to 250X250 pixels) of the thorax were segmented to 32 separate tissues,
requiring 36 hours of processing. Also other medical images including CT were analyzed accurately.
The applicability of Anatomatic for volumetric analysis of brain lesions was estimated. 23 MS patients (3D FSE
T2 MRI, 21 slices per patient) and 43 brain infarct patients (T2, 21 slices per patient) were segmented into MS
plaques and infarctions using the Anatomatic software. Segmentation time varied from 5 to 20 minutes for a patient
depending on the number of lesions and quality of images. The inter and intra-observer studies (in which six randomly
selected MS patients were segmented) demonstrated the 7% and 3.5% variability respectively. The volumetric accuracy
of the segmentation methods was demonstrated by segmenting fluid filled syringes. The test demonstrated in average
1.5% error between real and segmented volumes.
The multimodal visualization library and software (Medimag) was tested with varying test material, including
raw and segmented MR and CT images. The tests were performed on a 200MHz Pentium Pro based PC, equipped with a
24bit color display and 64MB of RAM. Surface reconstruction requires 20 seconds of processing (250X250X99 voxels),
but thereafter displaying and different projections require only one second. For a 512X512X100 voxel image the
preprocessing required about two minutes of processor time. The produced 3D images were excellent, the Medimag
software was easy to use. The software also enabled multimodal combination of medical images and measured or simulated
information.
The 3DEEG software was tested on a 200MHz Pentium based computer running WindowsNT. Surface reconstruction (250X250X100
voxels MRI), required 2 seconds of processing, definition of electrode locations and electrode mesh required 20
seconds. Displaying the electric field operated very fast, enabling screen updates with the frequency of 5 Hz (10-20
electrode system, 48 triangles). The updating speed is proportional to the number of triangles (100000 triangles,
5 seconds of processing). When the locations of computed electrode locations were compared to marked locations,
little variability was observed. Therefore the use of electrodes with MRI positive markers was recommended. 3DEEG
enables also the multimodal visualization of cortical EEG together with segmented brain images.
4. Discussion
The developed algorithms and software form an efficient and functional image analysis package for neurological
and radiological applications. They have been applied in clinical research projects concerning volumetric analysis,
reconstruction of resistive head models, and visualization of measured or simulated EEG signals. The emphasis of
the study has been put on functionality, intuitivity, efficiency, and versatility, rather than on the development
of new image processing principles.
The IARD segmentation algorithm is based on basic image processing techniques, such as image filtering, thresholding,
and region growing, but involves prior knowledge of brain anatomy; information (anatomical and radiological appearances)
of different neural structures are applied as the priority in integrating different tissues (decision trees) enabling
simultaneous detection of numerous tissues. Anatomatic and IARD algorithm were primarily implemented in order to
construct resistive head models. Due to relatively high resistivity of bone tissue, it was important to detect
the skull as accurately as possible. Therefore interactive approach was selected to ensure exact segmentation.
However, this feature requires more time, but errors caused by movement artifacts, anatomical inconsistencies or
pathologies, and automatic segmentation are corrected.
Numerous automatic segmentation techniques capable of segmenting the brain tissues have been introduced, based
on elementary segmentation techniques, statistical analysis, and neural networks. Such methods are able to detect
CSF, gray matter, and white matter, and possibly other tissues, like scalp-bone and brain lesions but not the skull.
The methods involve the use of multiple MR images, such as T1, T2, and PD images. Some techniques are capable in
segmenting brain structures from single MR images but require huge amount of processing (2 hours - 2 days). In
this point of view, IARD algorithm appears to be efficient and versatile. However, due to IARDs interactive character,
the borders of different tissues are defined manually leading to variability of segmentation results. E.g., the
separation of gray and white matter depends on the opinion of the person who controls the segmentation.
The versatility of Anatomatic was demonstrated by segmenting various different medical images [41]. Relatively
demanding task was to separate head MRI (T1) to 11 different tissues and VHM images to 32 tissues. Such procedures
cannot be implemented automatically, and on the other hand, conventional manual segmentation would be far too complex
and time demanding. Nevertheless, time usage was quite high also when IARD technique was applied (24 and 36 hours).
Anatomatic and IARD algorithm were also applied in volumetric analysis of brain lesions [42]. MS plaques and
infarcts were segmented from conventional MR images. In addition to the normal segmentation tools of the Anatomatic,
several manual editing techniques were developed, involving simple drawing tools and priorities [41]. The advantage
of the technique is to use three separate thresholds instead of one. Such presentation is capable of emphasizing
lesions of different intensities. However, the accuracy of the results depends on the skill of the person who controls
the segmentation. In order to estimate the effect of the operators, inter- and intra-observer studies concerning
MS plaques were carried out demonstrating 7% and 4% variability between observations, respectively. Taking into
account the difficult appearance of MS plaques the variabilities are relatively low. However, the operator must
be an expert on radiology and neurology. According to Van Waesberghe [45], in addition to the variability of segmentation,
different MR scanners can cause 1-2% variability, patient movement and reposition 5-10% variability, and too high
slice thickness 0.1-7.5% variability in volumetric studies. Therefore an accurate validation must be carried out
before serial clinical studies. Validation of the segmentation of brain tissues is very difficult; accurate non-invasive
estimation of shapes and volumes is impossible to complete. Although cadavers could be applied, only the brain
and skull can be measured accurately, but not CSF, gray, and white matter. Further, the shape and size of the brain
changes slightly after removing from skull.
The developed 3D visualization library and software (Medimag) proved to operate efficiently with raw and segmented
MR images. Its versatility enabled the integration of different objects (e.g., MRI data of the head and segmented
brain structures), different volume cuts, transparency, and multimodal visualization. Such properties enable efficient
representation of images aiming to medical utility; tumors can be presented in 3D together with surrounding MRI
data, and computed electric fields can be presented on the 3D surface or internally using volume cuts. The characteristics
of the library seem to be superior when compared to several other 3D libraries [43], due to performance and class
structure. However, state of the art techniques, like Ray Tracing and Radiosity are able to produce more photorealistic
images. Such techniques are not appropriate to medical routine due to demanding processing. The shading technique
based on gravity has not been applied before in 3D visualization packages. In order to increase the capabilities
of the library and software, the use of parallel computing, multimedia instruction sets, and 3D graphic accelerators
should be evaluated. Also the use of parametric surfaces would be interesting in future development.
The 3DEEG software together with brain mapping functions were implemented and tested successfully. The technique
involves Ray Casting, triangular facet surface between electrodes, virtual electrodes, and interpolation of EEG
potentials based on nearest electrodes [38]. Ray Casting and triangles have not been reported earlier among EEG
brain mapping methods, but parametric surfaces [35,37] have been applied. The processing requirements are modest
hence enabling very fast updating speed (6 Hz). Although 3DEEG operates efficiently, the image quality suffers
slightly when compared to parametric methods. In order to improve the properties of the 3DEEG, for instance better
interpolation technique should be possessed [35]. The other multimodal visualization technique capable of displaying
simulated electric fields applies triangular facet surface, Phong shading, and linear interpolation inside surface
elements. Due to the great number of surface elements (about 100000 triangles), linear interpolation operates efficiently
and the quality of produced images is high. Nevertheless, rendering speed is relatively fast requiring about 5
seconds for each new projection.
The three above mentioned techniques and software are important tools in order to construct neurologists workstation
software. At the moment, the software applies similar data formats enabling efficient data transfer. However, in
the future they should be able to use standard formats, like DICOM and EDF. In addition, it could be useful to
integrate some structures of the software, e.g., in order to display segmented structures in 3D during segmentation.
Also the integration of 3DEEG and Medimag software could be useful in improving the brain mapping image quality.
The segmentation capabilities will be improved in future development, not only because of different segmentation
techniques, but due to new MRI sequences. The developed software package could be developed further to study segmentation
applications in pathology, interventions, virtual operations, scopies, and cardiological applications. Because
the use of digital analysis of medical images is increasing rapidly in numerous research centers and in clinical
practise, the prospectives for segmentation and 3D visualization techniques are good. They will not replace conventional
image analysis techniques yet but are applied side by side. As soon as the segmentation techniques have been automated
and their accuracy and reproducibility are sufficient, these new techniques will improve diagnostic capabilities.
Segmentation has numerous clinical uses and therefore it is likely that in future most medical imaging devices
will include a segmentation application. The volumetric analysis will become a routine tool in radiology and reduces
the total costs; better diagnostic capabilities will decrease the costs caused by wrong diagnoses and inaccurate
follow-up of disease progression. Segmentation combined with anatomical atlas is likely to improve the diagnostic
accuracy of brain lesion studies.
Acknowledgements
The study was supported by Medical Research Fund of the Tampere University Hospital, Emil Aaltonen Foundation,
Wihuri Foundation, Ulla Tuominen Foundation, Instrumentarium Science Foundation, Foundation of Technology, Finnish
Cultural Foundation, and Ragnar Granit Foundation.
References
[1] Zhang Y (1997): Evaluation and comparison of different segmentation algorithms, Pattern Recognition
Letters, 18:963-974.
[2] Laarne P (1996): Solution of the electric forward problem using a realistic head models, Licentiate
thesis, Tampere University of Technology, Finland.
[3] Lehtinen M, Forsman K, Malmivuo J, Eskola H (1996): Effects of skull and scalp thickness on
EEG, Medical & Biological Engineering & Computing, 34(suppl. 1):263-264.
[4] Babiloni F, Babiloni C, Carducci F, Del Gaudio M, Onorati P, Urbano A (1997): A high resolution
EEG method based on the correction of the surface Laplacian estimate for the subjects variable scalp thickness,
Electroencephalography and Clinical Neurophysiology 103: 486-492.
[5] Yvert B, Bertrand O, Echallier J, Pernier J (1995): Improved forward EEG calculations using
local mesh refinement of realistic head geometries, Electroencephalography and Clinical Neurophysiology 95:381-392.
[6] Scherg M (1992): Functional imaging and localization of electromagnetic brain activity, Brain
Topography 5(2):103-111.
[7] Scherg M, Ebersole J (1993): Models of brain sources, Brain Topography 5(4):419-423.
[8] Scherg M, Ebersole J (1994): Brain source imaging of focal and multifocal epileptiform EEG activity,
Clinical Neurophysiology 24:51-60.
[9] Hombach V, Meier V, Burkhard M, Kuhn E, Kuster N (1996): The dependence of EM energy absorption
usin human head modeling at 900 MHz, IEEE Transactions on Microwave Theory and Techniques 44(10):1865-1873.
[10] Frank J, Stone L, Bash C, Calabresi P, DeCarli C, Maloni H, Stone R, McFarland H (1996): Is
there a relationship between contrast enhancing MRI lesions and change in bulk white matter in relapsing remitting
multiple sclerosis patients treated with beta 1b, Proceedings of the International Society of Magnetic Resonance
in Medicine:168.
[11] Filippi M, Yousry T, Horsfield M, Baratti C, Mammi S, Volz R, Comi G (1996): Quantitative assessment
of MRI lesion load in multiple sclerosis: A comparison of conventional spin-echo with fast-fluid attenuated inversion
recovery, Proceedings of the International Society of Magnetic Resonance in Medicine:541.
[12] Gawne-Cain M, Webb S, Tofts P, Miller D (1996): The effect of repositioning inaccuracies on
intracranial MS lesion volume measurements, Proceedings of the International Society of Magnetic Resonance in Medicine:543.
[13] Losseff N, Wang L, Lai H, Yoo D, Gawne-Cain M, McDonald W, Miller D, Thompson A (1996): Progressive
cerebral atrophy in multiple slcerosis, a serial MRI study, Brain 119:2009-2019.
[14] Wang L, Kilpatrick N, Lai H, Gawne-Cain M, Miller D (1996): A survey of the distribution of
lesion size in multiple sclerosis: Implication for the measurement of total lesion load, Proceedings of the International
Society of Magnetic Resonance in Medicine:546.
[15] Samarasekera S, Udupa J, Miki Y, Wei L, Grossman R (1997): A new computer-assisted method for
the quantification of enhancing lesions in multiple sclerosis, Journal of Computed Assisted Tomography, 21(1):145-151.
[16] Kamber M, Shinghal R (1995): Model-based 3-D segmentation of multiple sclerosis lesions in magnetic
resonance brain images, IEEE Transactions on Medical Imaging, 14(3):442-453.
[17] Johnston B, Atkins S, Mackiewich B (1996): Segmentation of multiple sclerosis lesions in intensity
corrected multispectral MRI, IEEE Transactions on Medical Imaging 15(2):154-169.
[18] Velthuizen R, Clarke L, Phuphanics S, Hall O, Bensaid A, Arrington J, Greenberg HM, Silbiger
M (1995): Unsupervised measurement of brain tumor volume on MR images, Journal of Magnetic Resonance Imaging 5:594-605.
[19] Peck D, Windham J, Emery L, Soltanian-Zedeh H, Hearshen D, Mikkelsen T (1996): Cerebral tumor
volume calculations using planimetric and eigenimage analysis, Medical Physics 23:2035-2042.
[20] Bryan R, Manolio T, Schertz L, Jungreis C, Poirier V, Elster A, Kronmal R (1994): A method of
using MR to evaluate the effect of cardiovascular diseases on the brain, American Journal of Neuroradiology 15(9):1625-1633.
[21] Mathews V, Barker P, Bryan R (1992): Magnetic resonance evaluation of stroke, Magnetic Resonance
8(4):245-263.
[22] Fox N, Freeborough P, Rossor M (1996): Visualizing and quantifying rates of atrophy in Alzheimers
disease from registered serial 3D MRI of the brain, Proceedings of the International Society of Magnetic Resonance
in Medicine:234.
[23] Freeborough P, Woods R, Fox N (1996): Accurate registration of serial 3D MR brain images and
its application to visualizing change in neurogenerative disorders, Journal of Computed Assisted Tomography 20(6):1012-1022.
[24] Zamora L, Jiang Z, Kadi A (1994): Computer-assisted neurosurgery system: Wayne State University
hardware and software configuration, Computerized Medical Imaging and Graphics 18(4):257-271.
[25] Gerke M, Vogt H, Fiekert W, Kretschmann H (1996): 3D reconstruction of neurofunctional structures
for neuroimaging, Computerized Medical Imaging and Graphics 20(1):1-13.
[26] Lorensen W, Cline H (1987): Marching cubes: A high resolution 3D surface construction algorithm,
Computer Graphics 21(4): 163-169.
[27] Watt A (1993): Computer graphics, second edition, Addison-Wesley Publishing Company 1992.
[28] Watt A, Watt M (1992): Advanced animation and rendering techniques, ACM Press 1992.
[29] Foley J, van Dam A, Feiner S (1994): Introduction to computer graphics, Addison-Wesley Publishing
Company, 1994.
[30] Santarelli M, Positano V, Landini L (1997): Real-time multimodal medical image processing: A
dynamic volume rendering application, IEEE Transactions on Information Technology in Biomedicine 1(3):171-178.
[31] George J, Aine C, Mosher J, Schmidt D, Ranken D, Schlitt H, Woods C, Lewine J, Sanders J, Belliveau
J (1995): Mapping function in the human brain with magnetoencephalography, anatomical magnetic resonance imaging,
and functional magnetic resonance imaging, Journal of Clinical Neurophysiology 12(5):406-431.
[32] Medina V, Hassainia F, Langevin F, Gaillard P (1994): Three dimensional representation of brain
electrical activity, Brain Topography 7(1):53-61.
[33] Lopes da Silva F (1993): Computer assisted EEG diagnosis: Pattern recognition and brain mapping.
In electroencephalography: Basic principles, clinical application and related field. Eds. E Niedermayer and F Lopes
da Silva, William and Wilkins, 1063-1086.
[34] Brinkman B, OBrien T, Robb R, Sharbrough F (1997): Localization, correlation, and visualization
of electroencephalographic surface electrodes and brain anatomy in epileptic studies, Medical Imaging 1997, SPIE
vol 3033:159-169.
[35] Perrin F, Pernier J, Bertrand O, Giard M, Echallier J (1987): Mapping of scalp potentials by
surface spline interpolation, Electroencephalography and Clinical Neurophysiology 66(1):75-81.
[36] Perrin F, Pernier J, Bertrand O, Echallier J (1989): Spherical splines for scalp potential and
current density mapping, Electroencephalography and Clinical Neurophysiology 72(2):184-187.
[137] Koles Z, Paranjape R (1988): Topographic mapping of the EEG: an examination of accuracy and
precision, Brain Topography 1(2):87-95.
[38] Heinonen T, Lahtinen A, Häkkinen V. Implementation of Three-Dimensional EEG Brain mapping,Computers
and Biomedical Research, in press.
[39] Pidoux B, Poirot F (1990): Tridimensional cartography of potentials recorded on the human scalp:
Interpolation with tridimensional functions, CR Acad Sci III 311(1):1-6.
[40] Heinonen, T., Eskola, H., Dastidar, P., Laarne, P. & Malmivuo, J. 1997. Segmentation of
T1 MR Scans for Reconstruction of Resistive Head Models. Computer Methods and Programs in Biomedicine 54, Ireland.
Elsevier Science Ltd.. p. 173-181.
{41] Heinonen, T., Dastidar, P., Kauppinen, P., Malmivuo, J. & Eskola, H. 1998. Semi-automatic
Tool for Segmentation and Volumetric Analysis of Medical Images. Medical & Biological Engineering & Computing
36, 3, p. 291-296.
[42] Heinonen T, Dastidar P, Eskola H, Frey H, Ryymin P, Laasonen E. 1998. Applicability of semi-automatic
segmentation for volumetric analysis of brain lesions, Journal of Medical Engineering & Technology, 33(4):
173-178.
[43] Heinonen T, Visala K, Blomqvist M, Eskola H, Frey H. 1998. 3D Visualization library for multimodal
medical images, Computerized Medical Imaging and Graphics, 22(4): 267-273.
[44] Dastidar P, Heinonen T, Vahvelainen T, Elovaara I, Eskola H. Computerized volumetric analysis
of lesions in multiple sclerosis using new semi-automatic segmentation software, Med. Biol. Eng. Comput, In press.
[45] van Waesberghe J, Filippi M, Gawne-Cain M, Nijeholt G, Barkhof F (1996): Reproducibility of
measuring MRI lesion load postmortem in multiple sclerosis: Influence of the scanners and of slice thickness, Proceedings
of the International Society of Magnetic Resonance in Medicine, pp. 532.
|

 Home
Current Issue
Table of Contents
Home
Current Issue
Table of Contents






